BrainSuite
MRI Segmentation, Analysis, and Visualization
BrainSuite is a collection of open source software tools that enable largely automated processing of magnetic resonance images (MRI) of the human brain. The major functionality of these tools is to extract and parameterize the inner and outer surfaces of the cerebral cortex, to segment and label gray and white matter structures, and to analyze diffusion imaging data. BrainSuite also provides several tools for visualizing and interacting with the data. BrainSuite is supported by NIH Grant R01-NS074980 (PIs: David Shattuck and Richard Leahy).
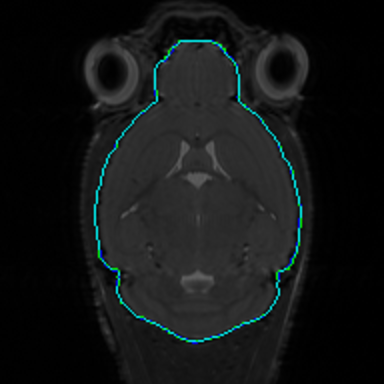
Preclinical Imaging
A Toolkit for Analysis and Visualization of Preclinical Rodent Neuroimaging Experiments
This project will develop an open-source suite of software for the processing, analysis, and visualization of rodent brain imaging data, which will enhance the ability of neuroscience researchers to perform preclinical studies on rodent models. This software will provide advanced capabilities for analyzing multimodal imaging data, including imagery from magnetic resonance imaging and optical microscopy studies, with the goal of improving understanding of processes and mechanisms underlying neurological diseases and disorders and their potential treatment. This project is supported by NIH Grant R01-NS121761 (PIs: David Shattuck and Allan MacKenzie-Graham).

Image Analysis of 3D Models of Idiopathic Pulmonary Fibrosis
Working with the research team led by Dr. Brigitte Gomperts, we are developing image analysis tools for high-throughput screening of an IPF model.
This project is supported by NIH Grant U01-1HL153000 (PIs: Brigitte Gomperts, John Belperio, and Robert Damoiseaux)

Virtual Reality
Multiuser virtual reality environment for visualizing neuroimaging data
We recently developed a virtual reality environment interactive visualization of neuroimaging data. Our system uses the HTC Vive to present a room-sized virtual experience for data exploration. Though still an early work, it offers a new way of exploring structural and diffusion MRI of the brain. In particular, our framework provides interactive visualization of volumetric MRI, surface models, tensor glyphs, and streamline tractography.
More information on this system can be found in our recent paper, “Multiuser Virtual Reality Environment for Visualizing Neuroimaging Data”.
More information on this system can be found in our recent paper, “Multiuser Virtual Reality Environment for Visualizing Neuroimaging Data”.
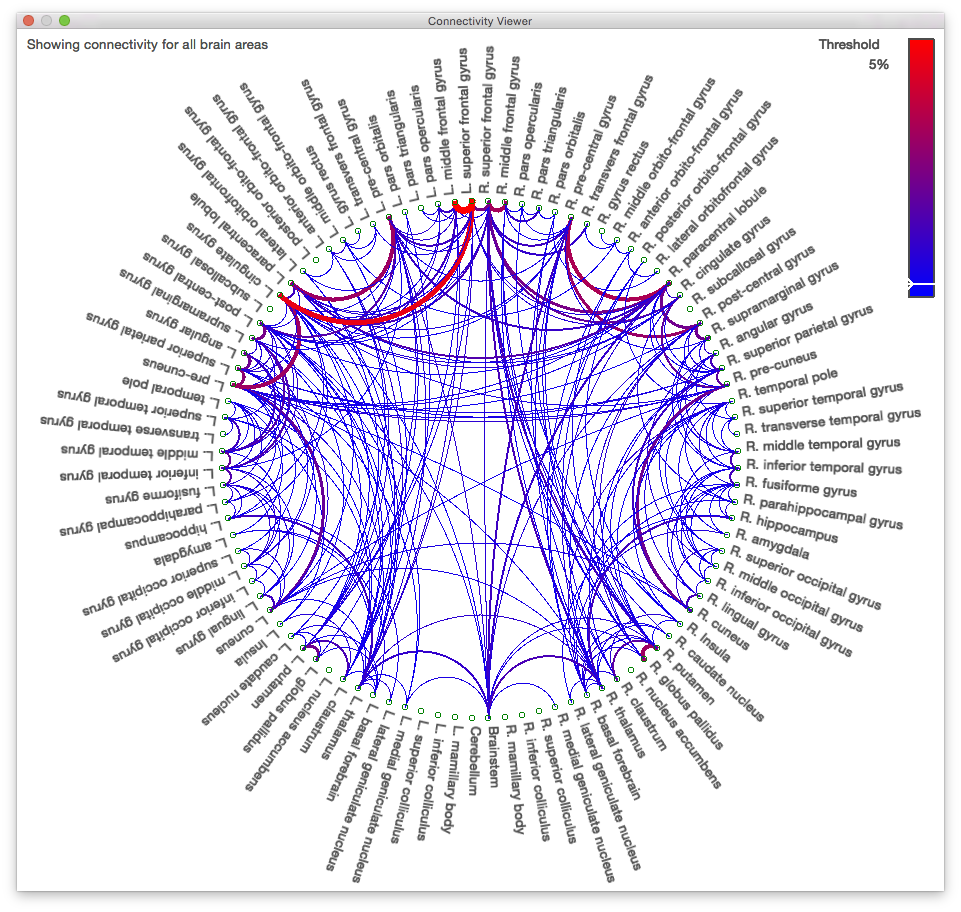
Perturbation of the Treatment Resistant Depression Connectome by Fast-acting Therapies
Connectomes Related to Human Disease
We are working with Dr. Katherine Narr and her research team on the Fast Acting Treatments of Depression project (NIH Grant U01 MH110008). This project will leverage optimized non-invasive MRI technologies and normative data available through the NIMH/NIA-funded Human Connectome Project to: 1) identify connectome-specific correlates and predictors of successful treatment outcome to 3 therapeutic interventions, each with a rapid onset of action; and 2) characterize alterations in neural connectivity associated with individual clinical, behavioral and physiological differences across treatment resistant depression.
Our group is working on the development of the image processing framework for this project.
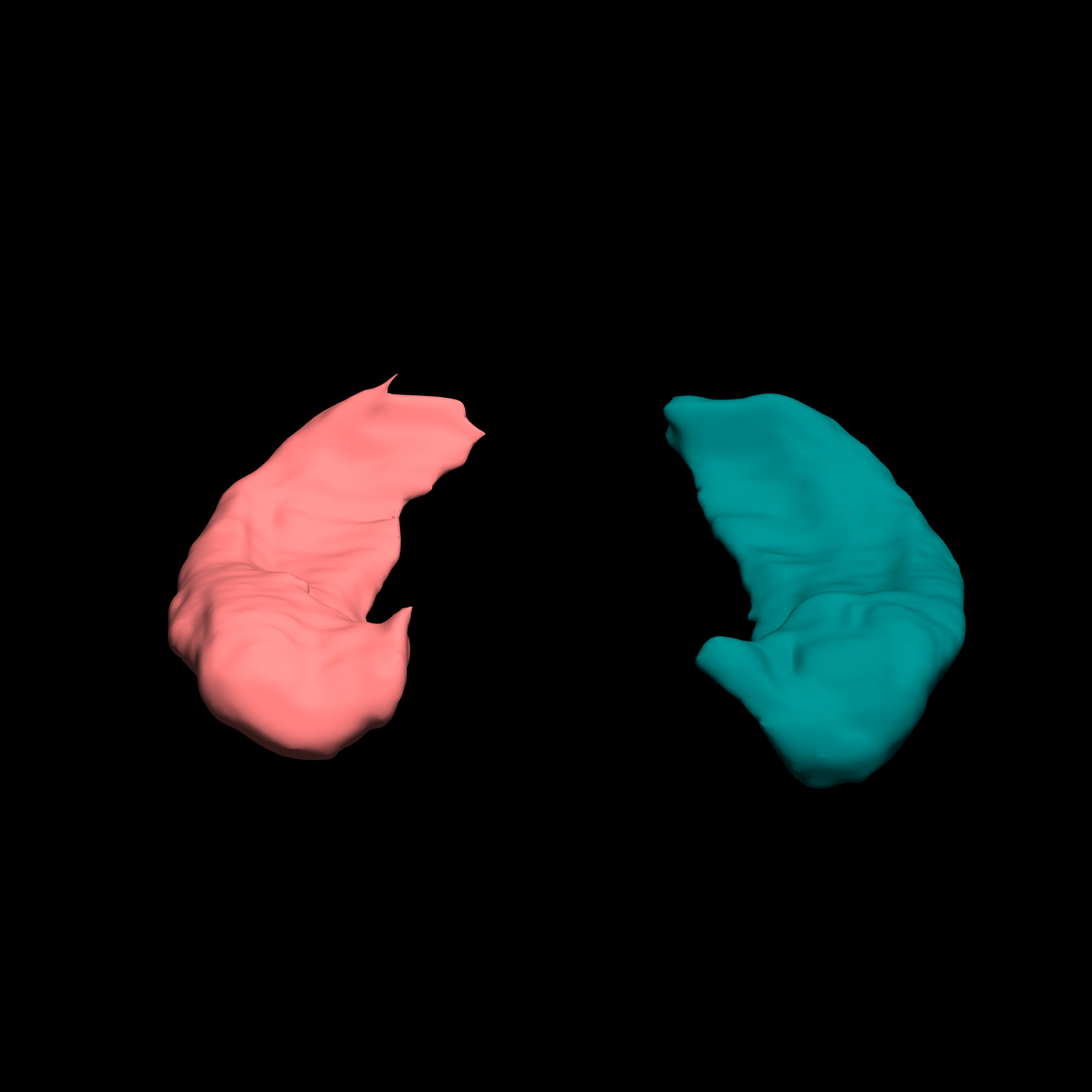

Hippocampal Sclerosis in Patients with Calcified NCC Characterizing
BRAIN Initiative project
We are working with Dr. Hector Garcia at the University Peruana Cayetano Heredia to analyze data from subjects affected by neurocysticercosis (NIH Grant R21 NS094976; PI: Garcia). Taenia solium neurocysticercosis (NCC) is a parasitic infection causing much neurological disease, particularly epilepsy, in most of the world. Old calcified brain parasitic NCC lesions are associated with damage to human hippocampal structures (hippocampal sclerosis, HS). The goal of this study is to assess a large existing cohort of individuals with calcified NCC to determine the proportion of them who have HS, the factors associated to HS, and to characterize the particularities of HS in NCC patients.
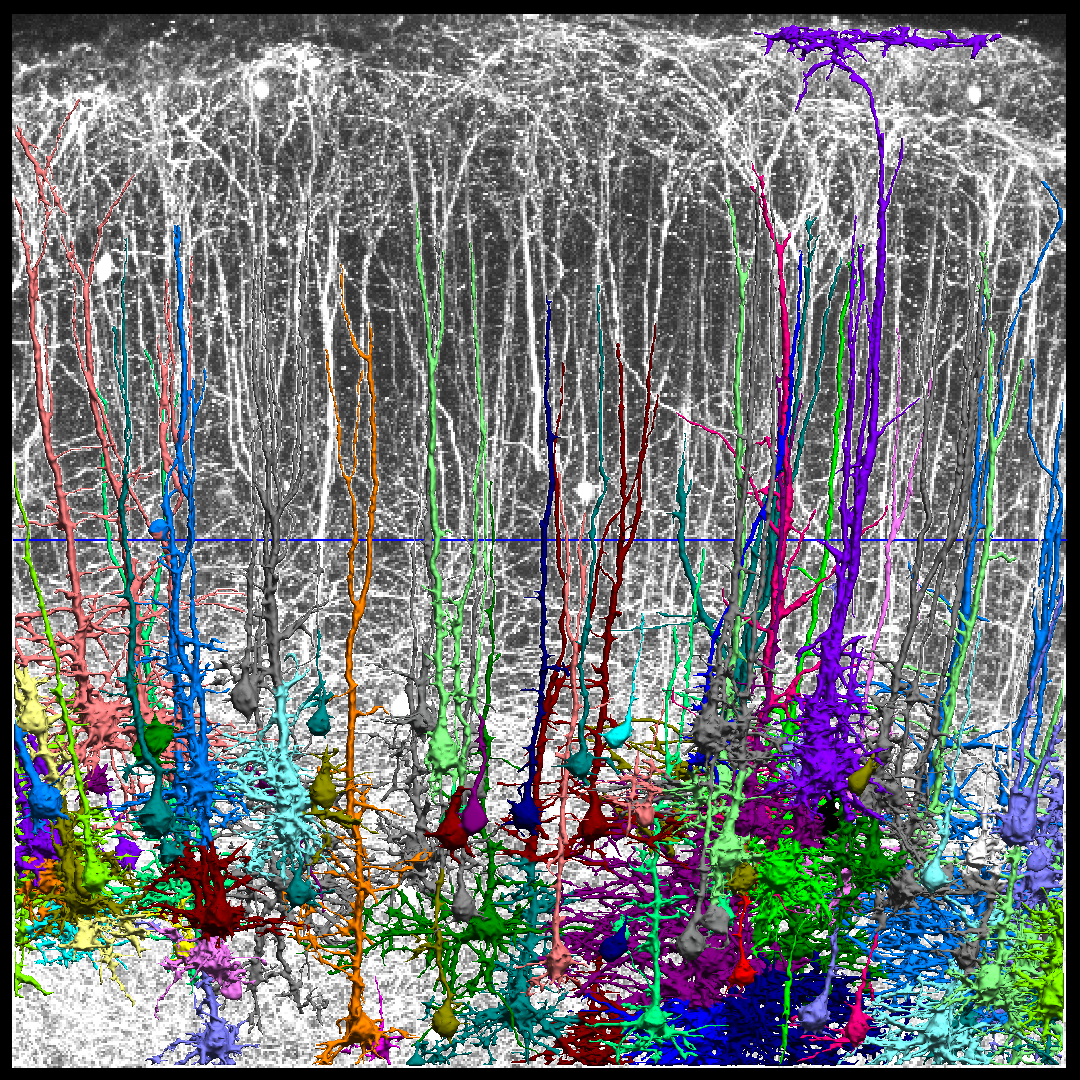
Defining Cell Types, Lineage and Connectivity in Developing Human Fetal Cortex
BRAIN Initiative project
We are working with Dr. Daniel Geschwind and his group team on developing new methods for analyzing fluorescence microscopy images of CLARITY data, funded by a BRAIN Initiative grant (NIH Grant U01 MH105991; PI: Geschwind). The goal of this project is to develop novel methods and tools to identify the individual cell classes that compose the human fetal cortex, determine the lineage of progenitor cell classes, and map the location, morphology, and connectivity of each cell class.